Generic implementation of dynamic neural fields
This Java framework serves two purposes : 1) Exploring and comparing the parameters or implementations of well studied dynamic neural fields (such DNF as the Continuous Neural Field Theory, aka CNFT) through a working Applet. 2) Letting the user combine existing classes or inherit from them to test and tune his own equations and algorithms (in a SDK like fashion). The Applet being used in this later case as a demonstration of the possibilities.
For direct manipulation of the Applet on this website, you must be able to run Java Applets in your web browser, but should not need to follow the installation procedure indicated below.
System requirements and download 
Requirements
Your system must satisfy the following requirements for the source code to compile or the program to run:
- Having the Java Runtime Environment (JRE6+) installed for the program to run. You can download it on the official Java website: http://www.java.com/en/download/
- To compile the source codes, having the Java SE Development Kit (JDK6+) installed. You can download it on the official Sun website: http://java.sun.com/javase/downloads/
Installation
For the installation of the various components required for the library to be functional, please refer the the original installation notes from the Java webpages or packages.
Click on the package icon on this page to download the SDK. You can either directly use it as is, or unzip the JAR file in a directory of your choice to edit and compile the file.
Once the components have been downloaded and installed, here are the possibilities to run the program:
- Double-click on the Jar file to start the program with the default arguments (if the .jar extension is associated with the correct program)
- Use the standard Java command lines in the file folder to run the program (
java -jar DNF.jarfollowed by any parameters of your choice) - Edit the files, compile them (
javac *.java) and run the program from the main class (java SimulAppletfollowed by any parameters of your choice) - Embed the Applet in a webpage and provide the adequate parameters (for instance the Applet embedded just below corresponds to the following syntax, although generated by quite different lines within the Wiki pages)
<applet code="SimulApplet.class"
codebase="../uploads/Software/DNFGeneric/"
archive="DNF.jar"
width="650" height="400">
<param name="scenario" value="asymmetry1" />
</applet>
If you get into errors, please check your classpath to see if all components are correctly registered, and consult the Java online help if needed.
Applet demonstration
This Applet illustrates how the DNF package can be used as a test platform, but also demonstrates the emergent properties of the CNFT (Continuous Neural Field Theory). It is also possible to change various parameters and the input dynamics as to observe their effect. For instance, the CNFT lateral connection profile, the integration constants, the number of stimuli and their trajectories or the level of noise can all be tuned. The kind of model used can also be changed from rate-coding to LIF spiking neurons (leak-integrate-and-fire).
Basic demonstration
With the default parameters used below, you should obtain an Applet with the following components:
- Input view (left): rendering of the input field that the CNFT will process
- Output view (right): rendering of the output neural field produced by the algorithm
- Time control panel: commands to control the simulation
The time control panel is itself composed of:
- Current time label: displays the current time in the simulation (expressed in seconds)
- Play/pause button: run the simulation indefinitely (>) or stop it (||)
- Step by step button: execute only one cycle through the update algorithm (|>)
- Target time text box: run the simulation until a particular time limit is reached (validated by pressing the return key)
- Reset button: reinitialize the output field (and hidden inner fields), as to restart the simulation (but the time value will not be put back to zero)
All neural fields can be manipulated with the mouse (all views are synchronized):
- Translate the views: move the mouse with the left button pressed
- Zoom in/out: respectively move the mouse right/left with the right button pressed
- Zoom in/out: as an alternative, use the central button (this can however interfere with the scrolling in some web browsers)
Although these pages are dedicated to the DNF package and not to the particular dynamics of the CNFT, you should observe the rapid convergence of the output field onto one of the stimuli, robustly tracking it despite the presence of the other stimulus. This should happen after about 9 seconds after a click on the play button, even though there is no difference in the strength of the two main stimuli.
Command line parameters
Additionally, a number of parameters can be specified by adding them at the end of the command line used to start the program (java -jar DNF.jar param1=value1 param2=value2) or within the applet configuration markup (see above):
gui:true,yesor1if you want to show the interface (default), else the computation will run in background at a much faster pace (useful for batch computations and scripting)interact:true,yesor1if you want to display the panel to edit the input field components and manage the statistics (data recorded over the simulation)views: determines the number of views to display in the interface (the level of detail and control you want to have on the intermediate computations)all: 4 views get displayed (top-left = input, top-right = weights, bottom-left = lateral interactions, bottom-right = output/focus)- (default): only the 2 views corresponding to the input and output fields
model: computational model used to update the output fieldspike: LIF spiking neurons implementation (leak-integrate-and-fire)- (default): rate-coding implementation
resolution: resolution of the field when using a matrix based implementations (which is the case for both the rate-coding and LIF implementations)scenario: dynamics of the inputs provided to the model. All stimuli have a 2D Gaussian profile with maximal intensity of 1.0 and follow centered circular trajectories.symmetry: 2 perfectly symmetrical inputsnoise: 2 identical inputs with a strong noise (uniform distribution with a 0.5 amplitude)distracters: 2 identical inputs and 5 distracters (taking random positions on the view)asymmetry2: 5 identical inputs with no noise (approximations break the symmetry)asymmetry1(default): 2 identical inputs with a low amount of noise (uniform with a 0.01 amplitude)
a1,s1,a2,s2: parameters of the lateral weight profile (w(s) = Ae-(s/a)² - Be-(s/b)²)alpha,tau,h: parameters of the differential equation governing the potential evolution (tau dup/dt = - up + sumk { w(||ap-ak||) uk }/alpha + ip + h))
Where the simulation of the previous section could correspond to (although the scenario parameter is not required as it takes the default value)...java -jar DNF.jar scenario=asymmetry1
... the command line for the simulation displayed below is:java -jar DNF.jar scenario=noise model=spike resolution=30
Changing the input parameters
When the program is launched with all the views and full graphical interface (interact=true views=all), the window should look like the one depicted below. It then includes the following components (some of them already presented earlier):
- Input view (upper left): rendering of the input field that the CNFT will process
- Weight view (upper right): rendering of the lateral connection weights (CNFT kernel)
- Competition view (lower left): rendering of the resulting lateral excitation/inhibition (convolution of the output with the weights)
- Output view (lower right): rendering of the output neural field produced by the algorithm
- Time control panel (bottom): commands to control the simulation
- Resolution control panel (1st right): matrix size used for computations (equivalent to
resolutionon the command line) - Noise control panel (2nd right): amount of noise added to the input signal (uniform noise by default)
- Distracters control panel (3rd right): number of randomly moving distracters added on the input signal (identical to the default Gaussian stimuli)
- Stimuli/CNFT control panel (4th right): selection of parameters for the input stimuli and CNFT profile (no stimulus selected by default, further described below)
- Statistics panel (5th right): panel to visualize and manage the recorded data over the course of the simulation (also described later)
Although changes on any of these parameters could be applied at anytime, it is better to pause the simulation first to avoid crashes (due to a bad implementation of the MVC model, for Internet optimization purpose)
To test the dynamics under different conditions, you can run the simulation with the default parameters in the above Applet (>), wait for a few seconds until convergence before stopping it (||), then change the amount of noise and/or distracters, and continue the simulation (>). You can also revert your changes and reset the simulation if you are unsatisfied with the results (for instance if your changes prevented the CNFT from correctly tracking one of the stimuli). For further details on the behavior of the standard CNFT using this Applet, you can check the CNFT demo page.
Changing the input stimuli
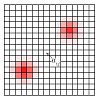
Following the advice shown in the panel labeled Gaussian parameters, you can left-click on any component of the input view to display and edit its parameters. A click in the view will select the closest stimulus. Further clicks at the same point will cycle through all overlapping stimuli (this also allows to clear the selection). Your selection will appear as a yellow circle that has no effect on the numerical simulation (see on the left view of Figure 1). It is possible to select and interact with components that do not appear on the views, because their intensity is currently to low, they are hidden by noise or the model has no yet been refreshed.
Here is a description of the components in the Gaussian parameters control panel:
- Add button: creates and select a new Gaussian component similar to the one previously selected (to add a new stimulus for instance)
- Remove button: remove the selected Gaussian component, as long as there remains at least one that can serve as a reference for creation (to actually remove all components through the interface, it is possible to set the intensity of the last Gaussian to zero, so it has no effect on the dynamics)
- Parameter trajectories: the following lines of components each correspond to a parameter of the Gaussian stimulus (x and y coordinates of the center of the Gaussian, standard deviation and maximal intensity), themselves described by a trajectory
The type of trajectory can be changed by left-clicking on the "<parameter> =" part of the interface. Be careful that the views might not be correctly refreshed in the web applet if the simulation is not running. The following types are implemented by default:
constant: the parameter will simply take a fixed value throughout the simulation (this is the default for the width and intensity of the stimuli, param = c)random: the parameter will take uniformly distributed random values centered around the first given value, with an amplitude defined by the second value (default for the position of distracters, param = c + a*random(-1,1))sinusoidal: the parameter will oscillate following a cosine function, the values respectively correspond to the center, amplitude, period in seconds and phase in [0,1] corresponding to [0,2PI] (position of the default stimuli, param = c + a*cos(2PI(t/period + phase)) with t being the current time of simulation in seconds)

Figure 1: Screenshot of the Gaussian parameter panel and associated selection on the views
All 3 types of trajectories are represented on Figure 2. Combining them can produce linear (although not of uniform speed of acceleration), elliptic, height-shaped and more complex movements when changing the center x and center y parameters, as well as contracting/expending or flashing stimuli when altering the intensity and width parameters. Notice that changes in the text boxes must be validated by pressing the enter key after modification to have an immediate impact on the input and model dynamics.
Changing the weight profile
Similarly to what can be done on the inputs, the CNFT weights can be dynamically updated. However, putting dynamical trajectories for the kernel definition will highly slow down the simulation (as optimization procedures won't be efficient anymore).
By repeatedly clicking in the weights panel, the excitatory and inhibitory components of the lateral connection profile can be selected (as it is a difference of Gaussians by default). You can then carefully alter the values, as specific small changes can have dramatic effects on the dynamics (for instance generating a self-reinforcing bubble of activity that will never disappear, or saturating the whole field with activity or inhibition). You can of course remove or add components on this field as well, but be prepared for the consequences...
Generated data

Figure 2: Screenshot of the statistics panel displaying the focus bubble position during the whole simulation (the straight line inside the circular trajectory correspond to the initial convergence period onto a stimulus)
For statistical analysis purpose, a set of variables are computed and recorder during the simulation:
Time: time in the simulation (in seconds)Focus x: x coordinate of the center of the bubble emerging on the focus/output field (between -0.5 and 0.5). As the position is computed as a center of mass on the whole neural map, the result is a rough approximation if too much noise is present, or meaningless if no bubble or several bubbles are presentFocus y: y coordinate of the center of the bubble emerging of the focus/output fieldInput x: x coordinate of the stimulus that is the closest to the focus bubble. This is not an evaluation as the statistical part of the program has access to the provided stimuli parameters before discretization occurs and noise is addedInput y: y coordinate of the stimulus that is the closest to the focus bubbleError distance: normalized distance between the closest stimulus and the focus bubble (the error does therefore not directly reflect the ability of the algorithm to keep track of the same stimuli)Likelihood: maximal activity on the focus field that reflects how much the focus interpretation of the sensory flow is reinforced (either by the input or by itself)Distortion: difference in shape between an ideal stereotyped bubble (based on the CNFT weight parameters) and the actual bubble present on the focus (the center approximation is used)Update time: time needed to make all the computations related to the update of the focus and competition fields (used for computational performance evaluation)
Please be aware that all these computations are only valid under a strong assumption that the dynamics should converge on a single bubble. Would the tested algorithm not be competition with global inhibition, the statistics should be totally different (and thus reimplemented in the SDK).
The two variables that should be displayed can be selected in the drop lists corresponding to the 2D axis of the graph. By default, the statistical panel displays the Error distance as a function of Time. Figure 2 however shows the emergent bubble trajectory (Focus x and Focus y selected) when the input is following a circular trajectory.
The graph can be cleared and the stored data reset by right-clicking on it. This does not reset the time in simulation, but allows to cut out the initial part of the data or focus on a particular event localized in time (as zooming in the views allows to focus on a specific spatial location).
Finally, if you downloaded the program and run it on your computer (or allowed the file access for the Applets and know where the default base web folder is located on your hard-drive), you will be able to save the simulation data as a CSV file[^CSV stands for Comma Separated Values. This is a readable text file where the first line corresponds to the variable names and following lines contain the raw data separated by comma characters.^] by hitting the Save statistics button. The default filename contains the date and time of the simulation, but you can change it in the field next to the button (the folder can however not be chosen in this version).
SDK structure and classes
Apart from its test bench status for DNF dynamics, the Applet can be considered as a collection of classes forming a SDK for the extension of existing, or the creation of new neural field dynamics. You can refer to Fig.3 for the UML class diagram of the SDK. The object-oriented development led to a hierarchy of classes, that take care of the definition and updating of neural fields, optimizing the computations, storing experimental data, rendering the fields or user interactions. The classes will be presented based on the group they belong to on the figure (one group per color).

Figure 3: UML diagram of the SDK classes used in the DNF applet. A few classes have been removed for simplification purpose, and also because most of them are still under development. The classes are colored based on their function.
The sub packages provide useful classes for the program/Applet interfacing (applet), for displaying multivariate functions and plotting data points (plot), or to optimize DNF parameters with genetic algorithms (genalg not displayed on the UML figure). However and since they are not necessary to understand and create neural field models, they will not be covered in this tutorial. You can in any case refer to the Javadoc or to the comments within the source code.
Top-level updatable objects (white)
Most objects in the SDK inherit from this class, as it defines the timeline neural fields and dynamical object can use to update their state. Although the only excerpt of the code will be presented for the other classes, the short Updatable.java source code will reproduced and commented here in its entirety.
public abstract class Updatable { /** Current time for this object */ double time = 0; /** Update period (in seconds) */ double dt = 0.1;
The time variable is not defined as static, which means that several instances can be updated on different timelines. Similarly, the non-static dt variable means that instances can be updated asynchronously. However and to simplify the analysis of the dynamics, instances are often updated with simulated synchronous parallelism, with larger update periods being multiples of the smallest one. For instance, setting dt=1 for the input signals while the model is updated with dt=0.1 means that the input will appear static for 10 model cycles before being updated.
Both variables are defined in seconds and can of course take values below the millisecond (0.001) for fine grained temporal models.
/** Update the parameters of the objects if needed */ public void update(double time_limit) { while (time<time_limit) { // Compute the new state compute(); // Update the time time += dt; } }
The update function is at the core of the SDK, as it guarantees the synchronization of the timelines for related objects. If the method is called on a set of instances with the same time_limit parameter, this method asserts that the time will fall after execution within the same update period.
/** Compute the new state after dt seconds */ public abstract void compute(); }
Nevertheless, all the intelligence is shifted to the compute method, the generic updating being generally hidden to the end-programmer, but for specific composition objects. It is this abstract method that must be carefully implemented for all dynamic objects to keep the coherence of the model for each timestep of dt seconds.
Maps and neural fields (gray)
Map: This class is the top-level class for manipulating neural fields. It should be identified by a unique string, associated with the name variable. If no name is provided to the constructor, an ID is automatically generated following the pattern map# where '#' is a number incremented for each new unnamed map.
It is also qualified by the wrap variable that determines if the underlying manifold should be bounded or toric. Bounded means that trying to access a value outside the defined bounds of the map will either return a predefined value (or rise an exception) while the toric mode (wrap=true) will map the coordinates into the bounds (using continuous modular arithmetics).
The get method is used to access the potential or variable considered over the field. This is not only useful for optimized matrix implementations, but also for avoiding discontinuities and easing visualization. Notice that the SDK is here limited to 1D/2D maps (1D being a degenerated case leading to non efficient computations).
/** Get the activity at point (x,y) * @param x x coordinate on the map * @param y y coordinate on the map * @return activity at (x,y) */ abstract public double get(double x, double y);
MatrixMap: This class directly inherits the Map class but specializes it for matrix-like computations. This means that the field values will be mapped onto a discrete and bounded 2D manifold. The matrix is defined by its resolution and the corresponding bounds in the continuous domain.
/** Bound of the matrix in the continuous space */ protected Rectangle2D.Double bounds; /** Resolution of the matrix (in rows/unit) */ protected double resolution; /** Deduced dimensions of the matrix */ protected int xs; protected int ys;
The default bounds are [-0.5,0.5]², corresponding to a call to new Rectangle2D.Double(0.5,0.5,1,1). The size of the manipulated matrix is then computed and updated using the following expressions:
xs = (int)(bounds.width*resolution); ys = (int)(bounds.height*resolution);
For instance, with the default bounds and a resolution set to 10 rows/unit, the cell top-left cell (0,0) in the matrix will correspond to the continuous coordinates [-0.5,-0.5], while the cell (4,4) will correspond to the center of the field of coordinates [0,0]. The activity on the field can then alternatively be obtained for any point with the integer coordinates in [0,xs][0,ys] with the discrete version of the get method.
/** Get the activity at point (ux,uy) * (warning: no boundary check is performed as to speed up the access) * @param ux ux coordinate on the matrix (in [0;xs]) * @param uy uy coordinate on the matrix (in [0;ys]) * @return activity at (x,y) */ abstract double get(int ux, int uy);
Finally, a number of methods are defined to manipulate the matrix fields efficiently (copy, clone, like, equals, equalsRough) and combine/compute on them using standard matrix based operations (convolve, scale, apply, aggregate). For the last two methods, interfaces are defined to encapsulate the scalars methods applied on the matrices.
/** Class representing a double function of several arguments */ public interface Function { /** Apply the function to the given parameters * @param params parameters of the function * @return image of these parameters */ public double apply(double... params); } /** Class representing an aggregation function */ public interface Aggregation extends Function { /** Constants to access the elements */ public static final int R = 0; // result public static final int X = 1; // x coordinate public static final int Y = 2; // y coordinate public static final int V = 3; // value at (x,y) }
A class implementing the Function might for instance compute the sum of the parameters (return params[0]+params[1]), while and Aggregation method would combine the inputs into the result (params[R]+=params[V]/xs/ys to compute the mean activity over the field).
DenseMatrixMap: This is the dense matrix implementation of MatrixMap, for which a Java array is used to store and manipulate the data. Other classes could be implemented, sparse matrices for instance, as to speed up computations for spiking models. A lot of code in this class is dedicated to optimized convolutions, using the SVD and FFT, respectively used to produce truncated Singular Value Decompositions and Fast Fourier Transforms of the kernel so as to speed up computations (making complexity fall from O(n4) to O(n^3) and O(n2 ln n)).
DenseMatrixConversion: This class bridges the gap between maps that have been defined by a continuous function (see NoiseMap or Stimulus classes below) and arrays on which complex computations can be easily performed. The generated field is then limited by the resolution of the underlying matrix, but can be used as a buffer to keep trace of past values. For instance, keeping random values over a period of time can be done by setting
(dt) to the desired value and constructing a DenseMatrixConversion map over a NoiseMap.
MultiMap: This map encapsulate a group of sub-maps as to perform operations on them and structure the computations. This may for instance be used to separate layers when modeling the cortical columns, or to create a hierarchy of stimuli when generating artificial inputs for the system. In pratice, the sub-maps are stored in a Java hash map for fast access and modification, the map names being the keys (the main reason for defining the names as unique identifiers, although the same name might be repeated at different places in the map hierarchy).
java.util.Map<String,Map> maps; public synchronized Map get(String m) {...} public synchronized Map put(Map map) {...} public synchronized Map remove(String name) {...}
The update method is here modified in order to update all the sub-maps before the encapsulating map. Cycles in the dependency graph are allowed as for successive calls to the method with the same time_limit parameter, the program will only enter the loop once (and thus not lead to an infinite recursion).
@Override public synchronized void update(double time_limit) { // Update the components and this map simultaneously while (time<time_limit) { double t = time+dt; // Update all the submaps for (Map m : maps.values()) { m.update(t); } // Update this map super.update(t); } }
CompositionMap: This short utility class simplifies the process of combining several maps using a simple operation. An abstract Composition class is thus introduced, as well as two implementations for finding the Sum and Max of all sub-maps activities at each point of the field.
A full example of using such maps to define complex neural fields made of several layers and inputs composed of many stimuli is provided with the green CNFT classes described below.
Noise maps (pink)
NoiseMap: This is used for the generation of random noise, for example when testing the robustness of the neural fields or when trying to make a stable bubble of activity emerge whatever the input. This class does not store randomly generated values and a DenseMatrixConversion should therefore be used to buffer them.
The noise is characterized by its distribution with the noise variable, which is an instance of the Noise class. Two implementations are provided for the noise generation: UniformNoise (in [0,1]) and GaussianNoise (of standard deviation 1). The noise can then be amplified or diminished by adjusting the amplitude variable in the NoiseMap class.
Stimulus maps (blue)
Stimulus: This map was introduced to define activity distributions that should be dynamically parametrized. Local input stimuli can for instance follow predefined trajectories and be later combined into a single input activity map using the CompositionMap class. Similarly to the sub-maps in the MultiMap class, the set of parameters are unconstrained, identified by a name, stored in another hash map and can be accessed and modified using the methods introduced in the source code below.
HashMap<String,Parameter> params; public Parameter getParam(String p) {...} public Parameter setParam(Parameter param) {...} public double getValue(String p) {...}
All stimuli should at least define two parameters for their center coordinates, as they should be local and the user should be able to interact with them (select, modify or delete them). An abstract method named getCenter must therefore be implemented by all subclasses.
Although it was not specified earlier, all maps have access to a method to find the closest stimulus to a given point. This is particularly useful when dealing with complex sets of maps, as the user might want to select the stimulus closest to her mouse click, whatever the depth of the object within the map hierarchy. The calls to the getStimulus method are therefore propagated down the hierarchy, and results are combined when going back up. This method uses the getDistance(x,y) and abstract getShape() methods to determine if the requested coordinates lies within or outside the stimulus, and how far from its center.
Here again as in MultiMap, the overridden update method makes sure that all parameters are updated before updating the stimulus.
Gaussian: This class provides a particular implementation and use of the parameters so as to define a Gaussian distribution of activity over the field. This particular class of stimuli is characterized by four parameters: the center coordinates (Center x,Center y), a standard deviation (Width) and an amplitude (Intensity). In this case, the center of the stimulus of course corresponds to the center of the Gaussian profile, and a circle is returned for the shape.
To give an example of how a map activity can be explicitly defined in a continuous way using the get method, the source code for the Gaussian class is provided below. Notice that the wrap variable is tested to determine how the activity should be distributed (this is also to warn the end-developer/user that the bounds should be corrected as they are incorrectly fixed).
@Override public double get(double x, double y) { // Put in (x,y) the distance vector to the center x = abs(x-getValue(CENTER_X)); y = abs(y-getValue(CENTER_Y)); // Checking the wrapping slows the generation down // (check if the center is further than half the map size) if (wrap) { if (x>0.5) x = 1-x; if (y>0.5) y = 1-y; } double w = getValue(WIDTH); return getValue(INTENSITY)*exp(-(x*x+y*y)/(w*w)); }
Parameter: Parameters is a wrapper class for a Trajectory, simply enriching it with a name field to unambiguously identify the parameter. It thus requests an update of the trajectory every time the parameter value should be updated.
Trajectory: This abstract class simply introduces the abstract getValue method used to obtain a new value on this trajectory based on the current time (on the 'local' timeline).
Although this is not covered here as it is unimportant for the direct creation and manipulation of neural fields, you might notice that a parse method is provided in these classes so as to easily define the parameters from a command line or script.
All implementing classes correspond to different trajectories. Whereas the ConstTraj simply returns a constant value over time (that can however be modified by the program or user), the generic CompTraj class combines several ConstTraj with a function to be specified. Implementations include RandTraj (random values uniformly distributed in a ball specified by a center and radius) and CosTraj (values following a cosine trajectory defined by a mean value, amplitude, period and phase). The compute function for the second implementation is given.
@Override public void compute() { value = params[0].getValue()+params[1].getValue() *cos(2*PI*(time/params[2].getValue()+params[3].getValue())); }
Data management (yellow)
Profiling: This is a simple utility class to evaluate the execution performance of the models. The COMPUTER, SYSTEM, USER or CPU times can be obtained for any portion of your code, by calling the profile method. The time resolution however depends on your virtual machine and operating system. For instance in CNFT.java, you can find the following lines:
@Override public void update(final double time_limit) { update_time = Profiling.profile(new Runnable() { public void run() { CNFT.super.update(time_limit); } }, Profiling.USER); }
Statistics: This class stores and/or computes any data worth saving for the current model. It should therefore be redefined for any profound change to the neural field model currently evaluated. Nevertheless for similar models, this class can be directly exploited to compare their performance (computation speed, accuracy for tracking, convergence speed, robustness...). The plot package provide classes to store (WTrace) and plot (Plotter) any set of variables during the simulation.
For the CNFT models provided as an example (and manipulated in the Applet), the class has the following structure:
/** Constructor */ public Statistics(CNFT model) { super(new String[]{ INPUT_X,INPUT_Y,FOCUS_X,FOCUS_Y, TIME,ERROR_DISTANCE,LIKELIHOOD, UPDATE_TIME, DISTORTION }); this.model = model; }
This constructors defines the variable names (defined as constant character strings) that should be stored. The main method named update then computes these variables every time it is called and adds them to the stored trace. The variables might be obtained through more or less complex computations (center of mass of the focus, distance to the closest stimulus, distortion of the bubble relatively to an ideal bubble...)
/** Compute statistics on the CNFT */ public void update() { // Retrieve the focus field of the model (output) MatrixMap focus = (MatrixMap)model.get(model.FOCUS); // Estimated focus position double fx = ... double fy = ... // Likelihood of the result (= maximal value) double likelihood = focus.aggregate( -Double.MAX_VALUE, new MatrixMap.Aggregation() { public double apply(double... params) { return Math.max(params[R],params[V]); } } ); // Error relative to the selected input Gaussian (distance) double error = 1; // arbitrary maximal value double ic[] = {0,0}; // arbitrary center // Find the closest input Stimulus stimulus = model.get(model.INPUT).get(model.TRACK).getStimulus(null,fx,fy); if (stimulus!=null) { ic = stimulus.getCenter(); error = stimulus.getDistance(fx,fy)/Math.sqrt(2); // normalized distance } // Distortion of the output bubble double distortion = ...
Or they might be directly copied from the model/simulation (here the model.time or model.update_time variables updated in the CNFT instance). In the end however, all variables must be stored as the current 'state' of the simulation.
// Output the statistics as a multidimensional trace add(ic[0],ic[1],fx,fy,model.time,error,likelihood,model.update_time,distortion); }
User interface (orange)
The user interface elements are quite spread over the classes, thus partially violating an MVC architecture pattern. However, most methods relating to the Swing interface and rendering are named getSwingView or getSwingController in the Map, Stimulus, Parameter, Trajectory or Statistics classes (the method names should be made homogeneous). In the map class, the method code is the following:
public JComponent getSwingView() { return new SwingMap(this); }
In Statistics.java, the getSwingView method returns a Swing component for the real-time rendering of the data (as a 2D graph where you can select the plotted variables). It additionally gives the opportunity to save the data as a standard CSV file (Comma Separated Variables) through the graphical interface.
SwingView: This class renders any field as a colored image (negative activities in blue, and positive in red). The code will not be detailed here, as it is rather technical and uninteresting for the scientific aspects of the SDK. It registers StimulusListeners to handle events based on the user interactions through the GUI (stimulus selection events based on mouse clicks).
SimulAppet: This huge class deals with the interpretation of parameters from the command line (or from the Applet context), the setting up of the neural field model, the rendering of the neural maps, the interactions with the user to add/modify/remove stimuli or to modify global parameters (like the level of noise, number of distracters or resolution of the maps). Most importantly, it controls the update flow for the Updatable objects through the introduction of a time panel.
This class should probably be modified for major changes in the underlying neural field model, but can be used without much modification in most cases when just extending/modifying the DNF equations or replacing the CNFT by a similar model.
If you don't need user interactions and real-time rendering, and simply want to simulate the DNF dynamics as fast as possible, you can refer to the GATracking class present in the downloaded package. It is not represented on the UML diagram as its deals with the manipulation of the classes at a meta level to optimize the DNF parameters, but it provides the basic methods to run a simulation without having to apply any change in SimulApplet.java. Please check the genoToPheno method to see how to create and configure a CNFT model (or any other model of your choice derived from the green classes), as well as the simulAndEval method to control the execution of the model and retrieve data.
Sample implementation (green)
CNFT: The classes in this group are detailed here as they provide all the basics for defining and combining neural field within this SDK. This abstract class already include most ingredients to dynamically update a DNF activity in response to an external stimulation. The source code is extensively commented below.
The main map of the model is constructed as a MultiMap that will be able to update and compute its member maps. At this level, only two maps are defined and added to the hash map: the Inputs and the CNFT_Weights.
public CNFT(Map inputs, Map weights) { super("CNFT model"); if (inputs==null) inputs = getDefaultInputs(2,0,new UniformNoise(),0.25); put(inputs); if (weights==null) weights = getCNFTWeights(1.25,0.1,-0.7,1); put(weights); }
The Inputs are themselves defined as a CompositionMap adding the influence from Trackable stimuli, Distracters and Noise. In turn, the input stimuli (s1...sn) are defined as a CompositionMap of Gaussians that follow a periodic Trajectory, thus the use of CosTraj for the Center x and Center y parameters.
/** Configure the inputs */ public Map getDefaultInputs(int input_nb, int distracter_nb, Noise noise_distrib, double noise_amplitude) { // Trackable objects on the input map MultiMap track = new CompositionMap(TRACK); for (int s=0; s<input_nb; s++) { // Phase changes between inputs track.put(new Gaussian( "s"+s, new CosTraj(0,0.3,36,s/(double)input_nb+0), new CosTraj(0,0.3,36,s/(double)input_nb+0.25), new ConstTraj(stimulus_width), new ConstTraj(stimulus_intensity) )); } // Distracters on the input map MultiMap distr = new CompositionMap(DISTR); for (int d=0; d<distracter_nb; d++) { distr.put(getDefaultDistracter()); } // Additional noise on the input Map noise = new NoiseMap(NOISE,noise_distrib,noise_amplitude); // Configure the input map (trackable objects+distracters+noise) return new CompositionMap(INPUT,track,distr,noise); }
On the contrary, the Distracters (d1...dn) by default follow random trajectories (RandTraj for the center coordinates).
/** Get a new default distracter */ public Map getDefaultDistracter() { // Next distracter unique identifier distracter_id++; // Return the distracter return new Gaussian( "d"+distracter_id, new RandTraj(0,0.5), // random x in [-0.5;0.5] new RandTraj(0,0.5), // random y in [-0.5;0.5] new ConstTraj(stimulus_width), new ConstTraj(stimulus_intensity) ); }
All implemented CNFT models also share the lateral connectivity using a difference of Gaussian kernel. Such kernel is also defined as a map, that the composition (computing a Sum by default) of a local Excitation part (a Gaussian with a positive coefficient A) and a large Inhibition part (B<0 and a<b).
/** Configure the CNFT weights */ public Map getCNFTWeights (double A, double a, double B, double b) { return getCNFTWeights(new Gaussian(EXCITATION,0,0,a,A), new Gaussian(INHIBITION,0,0,b,B)); } /** Configure the CNFT weights */ public Map getCNFTWeights(Gaussian excitation, Gaussian inhibition) { return new CompositionMap(CNFT_WEIGHTS,excitation,inhibition); }
DenseCNFT: This class additionally specifies how the model should be transformed to handle matrices. The only method defined encapsulates the continuous maps into DenseMatrixConversion maps with the desired resolution. It also adds two new maps that will be used for intermediate (CNFT for the lateral competition term) and output computations (Focus).
public void setResolution(double resolution) { // Replace the input and cnft_weights maps by equivalent matrices put(new DenseMatrixConversion(get(INPUT),resolution)); put(new DenseMatrixConversion(get(CNFT_WEIGHTS),resolution)); // Configure the CNFT and focus maps put(new DenseMatrixMap(CNFT,resolution)); put(new DenseMatrixMap(FOCUS,resolution)); }
All together, these classes produce the map hierarchy illustrated on Fig.4. For each map, the class indicated in the lower right corner (but for CNFT model) corresponds to the sub-classes used to combine or define the map dynamics.

Figure 4: Sample maps hierarchy for the dense CNFT model.
DiscreteCNFT (named like this for strange historical purpose in my work): This class finally instantiates the CNFT equation in its rate coding version (as introduced by Amari in his seminal paper of 1977). The overridden update method specifies how the different fields should be updated. Although the hierarchies for the Inputs and CNFT_Weights are automatically updated, the CNFT and Focus field activity must be manually defined with the ordinary differential equation (noted ODE on the figure). The same of course holds true for the SparseCNFT class that implements a spiking neuron version of the CNFT.
public void compute() { DenseMatrixMap focus = (DenseMatrixMap)get(FOCUS); DenseMatrixMap cnft = (DenseMatrixMap)get(CNFT); // Compute the CNFT focus.convolve(get(CNFT_WEIGHTS),cnft); cnft.scale(1.0/(cnft.xs*cnft.ys)*(40*40)/alpha); // should be in the weights definition // Focus computation focus.apply(new DenseMatrixMap.Function() { public double apply(double... params) { return applyCNFT(params[0],params[1],params[2]); } private double applyCNFT(double input, double cnft, double focus) { return Math.max(0,focus + dt/tau*(-focus + cnft + input + h)); } },get(INPUT),cnft,focus); }
Thank you for reading this Applet/SDK tutorial, and thanks to all the CORTEX project members who helped or advised me before, during and after the development!
[^#^]